-
New Features
Ability to get correlated crosshair cursors in stacked spectra.
- It is now possible to get correlated cursors in stacked mode

New Data Analysis algorithms (Max Peak Position and Pick Alignment shifts).
- We have implemented two new algorithms to find the Maximum of the peak chemical shift and another by using the cross correlation (alignment) feature

Capability to export and import the Data Analysis table

Capability to export and import the Data Analysis table Capability to disable points for Data Analysis directly from the arrayed dataset
- It is now possible disable points directly from your arrayed dataset. You will notice how the applicable points in the XY graph will disappear and the signals on the stack plot will be highlighted in grey.
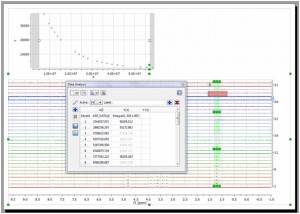
Capability to change the top and bottom margins of the stack plot.
- This is is really useful in to avoid clipping a stacked plot at the spectrum border. With this property you will be able to adjust the distance between the top of the spectrum object and the baseline of the uppermost spectrum.

Capability to export assignments info in journal format
- It is now possible to export your assignments by scripting
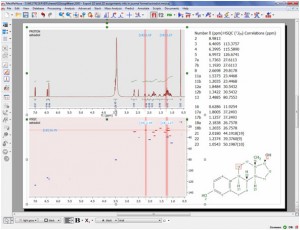
Manual Phase Correction pivot pointer is now easier to handle
- Just drag&drop the red little square to move the pivot pointer
 Capability to load/save spin systems from the Spin Simulation table.
Capability to load/save spin systems from the Spin Simulation table.- The user will be able to import XML files to synthesize 1D NMR spectra
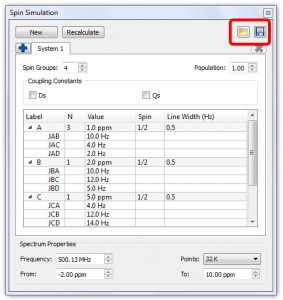
New icons in the Stacked NMR toolbar
- Now it is possible to increase/decrease the intensity of the traces from the toolbar.
Bugs Fixed
- Problems opening a NMR dataset containing a wrong mol file
- Difficulties opening NMR files in Jeol gxd and gxp format
- Mnova showed 2D plots importing some Bruker 2rr arrays
- Problems import phase correction parameters from stack datasets
- Problems setting up the Peaks Report from Script menu
- Problems running twice the GSD Spectra script from Export ASCII scripts menu
- Problems saving as SD file when molecules and spectra are opened
- Problems correcting baseline of R-M data with Bernstein Polynomial method
- NMR Peaks & Snap did not paste annotations
- Problems reading Lowest Frequency parameter in .dx files
- Problems with baseline of spectra from a document generated with Mnova-6.0.2
-
New Shimadzu and NetCDF ANDI-MS converters
- It is now possible to load Shimadzu and NetCDF datasets into Mnova MS
Bugs Fixed
- Problems opening big mass datasets from Bruker Daltonics
- Problems getting back to full spectrum after zooming too much
- Problems opening CG files from Agilent Chemstation without MS
- Problems trying to open ms datasets in MassLynx format
- Problems opening Bruker Xmass files without selecting the type from the Open dialog
- Wrong scale in some Bruker XMass datasets
What’s new in Mnova 6.2.1
0
Share.


