Mnova 14.3 has just been launched and includes new products, some plugins updates, a number of new features for many of our products, along with the usual bunch of bug fixes. You can read our new features overview on our changelog page. Here though, we would like to highlight what we consider the top changes and additions for our customers.
Download Mnova 14.3
You can read a brief description of the top highlights for Mnova 14.3 we've selected, in no particular order, below:
1. New Product! Mnova Screen 2D
Efficient Batch Processing Tools for Lead Discovery using Protein-Observed 2D NMR
This product is related to Mnova Screen, which can be used to process ligand-observed 1D NMR spectra (STD, T1rho, CPMG, WaterLogsy, etc.). Screen 2D processes the protein-observed 2D 1H-15N, 1H-13C, or H1-13C/15N dual heteronuclear correlation (HSQC or HMQC) spectra to find binding ligands based on chemical shift perturbations.
Read more about Mnova Screen 2D
2. New Product! Mnova Gears Chrom Cal
Automated Evaluation and Generation of Calibration Curves for Multiple Compounds Simultaneously
Generating well-defined calibration curves is crucial for accurate quantitation via chromatographic methods. With Chrom Cal, this error-prone and time-consuming process can be fully automated and streamlined for multiple compounds at the same time!
Read more here Mnova Gears Chrom Cal
3. New Product! Mnova Gears Chrom Reaction Optimization
Automated Software Solution for Chemical Reaction Optimization
With this new solution from Mnova Gears, determining ideal reaction conditions becomes an easy, hassle-free, and quick operation. Chrom Reaction Optimization allows you to track a defined set of chemical species across multiple parallel reactions to compare their levels and determine the best experimental conditions to employ.
Read more here Mnova Gears Reaction Optimization
4. Support of Waters Empower and Thermo Chromeleon files - MSChrom
We are delighted to tell you that Mnova MSChrom (formerly Mnova MS) can now import and handle Water Empower and Thermo Chromeleon files. This tool has been added to the Mnova Tools ribbon under the ‘Import’ dropdown menu. Note that this tool needs to have Empower and/or Chromeleon installed on the same computer to work.
This tool actually works via a two-step action: it exports data from Empower or Chromeleon to a file, and then imports that file into Mnova.
5. Support for Fractions (Waters MassLynx so far) - MSChrom
We have implemented a new converter to deal with fractions. It automatically imports the fraction information from a file while loading an MS file or folder. In this way, you can set up the fraction file name by provider in the Fraction sheet of the Mass Preferences import sheet.
You can also define rules to set the fraction's file name in the MS metadata function. Please note that only Waters-MassLynx fractions information is supported at this stage, but support will be extended to other fractions soon.
The information from the fraction file is shown in the Chromatogram Fractions table. A color is automatically assigned to each fraction.
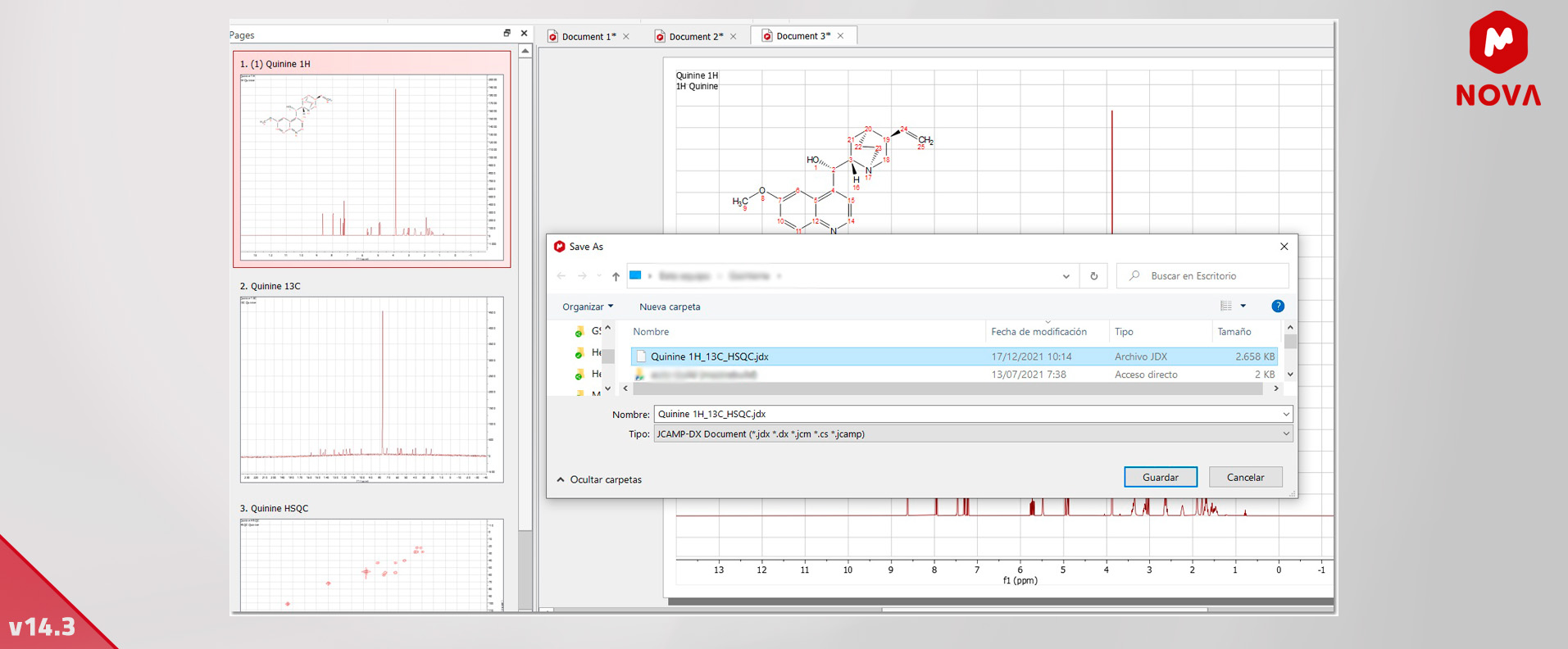
6. Save the Whole Document as JCAMP - Mnova General
We are pleased to announce that we have enhanced the way we handle JCAMP files. We have implemented a new file filter, "JCAMP-DX Document” (*.jdx *.dx *.jcm *.cs *.jcamp), which is analogous to the other JCAMP-DX except for the fact that when saving, it saves the entire document.
This converter will be always enabled no matter the selection made. Basically, you can save the entire document with several pages at the same time as a JCAMP file.
7. Improved JCAMP Export - NMR
We have improved a number of features to save the original settings when exporting as a JCAMP file, such as:
- 1D or 2D cuts
- Zoom
- Stack mode
- Traces in the spectrum
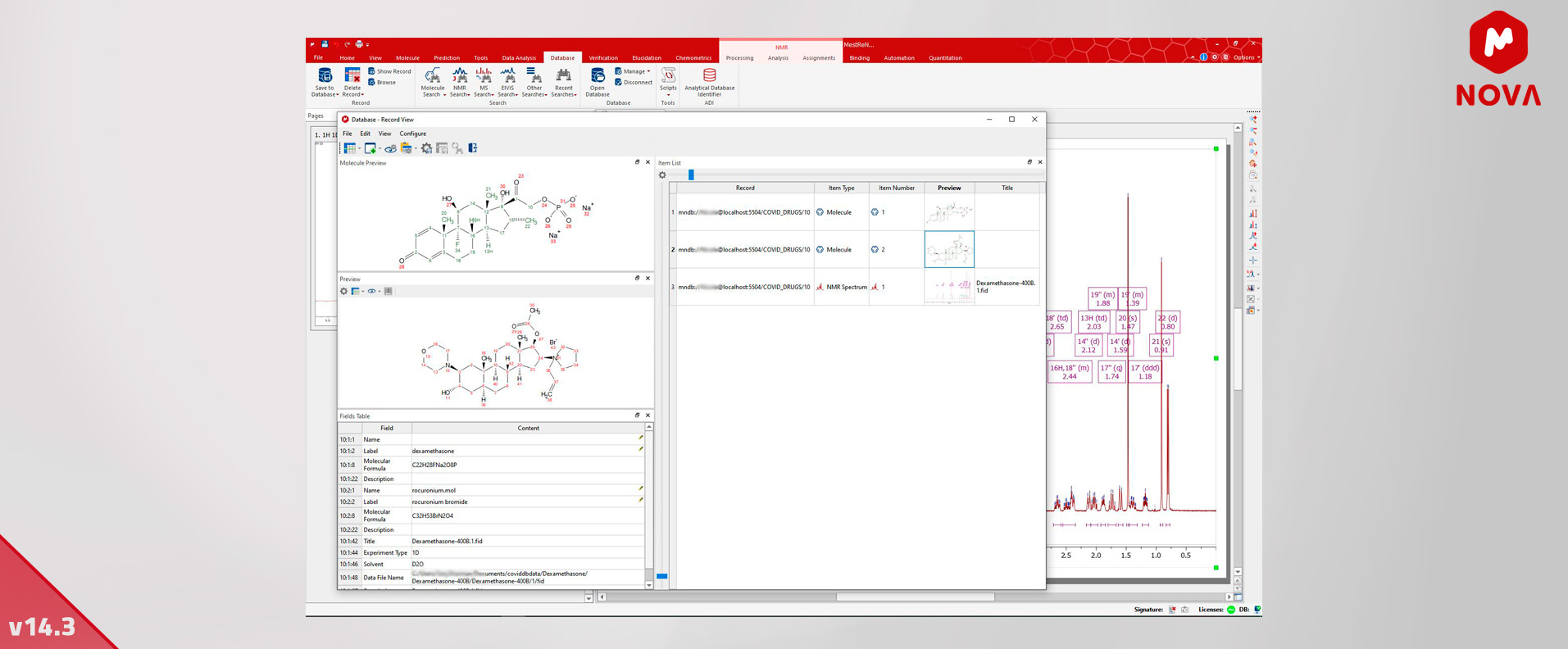
8. Allow multiple molecules in one DB record - Mnova DB
Mnova DB allows now to save several molecules in one record and the creation of databases without molecules. (The Molecule method is not mandatory anymore for a new database).
9. New Custom Mode to Select MS Spectra from Chromatograms - MSChrom
We have implemented a new crosshair mode, “Customized Peak”.
This mode can be used to select a given peak in a chromatogram, integrate a percentage of the MS scans of this peak, and automatically subtract scans before/after this peak. The implementation ensures that the integration has at least one MS scan. This mode has been added to the Mass section Tools > Crosshair/Select button menu. The corresponding scripting function has also been implemented.
You can also change the settings using the ‘Customized Peak Settings’ drop-down menu. In this way, the integration percentage can be modified, and the integration of the reference peak can be chosen.
10. System Suitability Calculations for Chromatographic Data - MSChrom
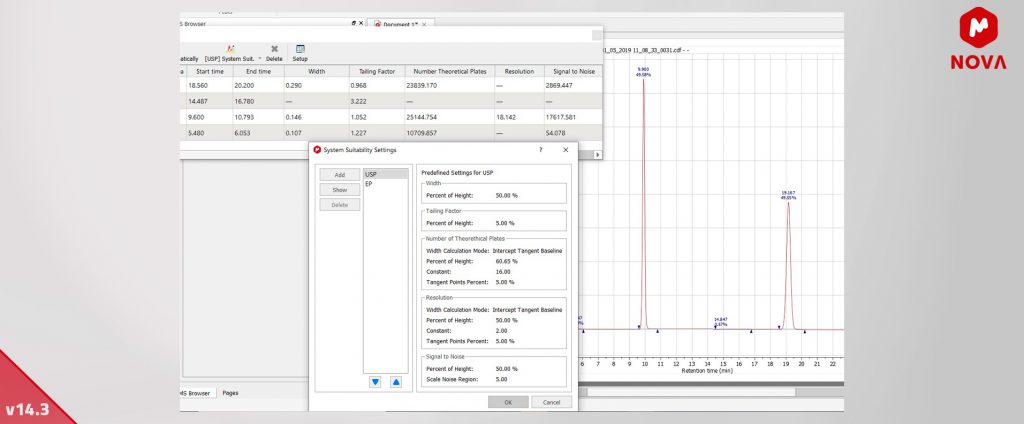
We have implemented system suitability calculations (width, resolution, number of theoretical plates, tailing factor and signal-to-noise) in the MSChrom plugin.
The first one will show the width of each peak at a specific percent of height. The other four remaining calculations will show the corresponding system suitability calculations for each peak.
A new button, "System Suit", has been added to the Chrom Peaks table toolbar. There are two predefined settings, USP and EP, which contain the values of the appropriate parameters for US and European Pharmacopoeia, respectively.
11. SIMCA Model Classification - Chemometrics
Soft independent modelling by class analogy (SIMCA) is a statistical method for supervised classification of data.
In order to build the classification models, the samples belonging to each class need to be analyzed using principal component analysis (PCA), from which only the significant components are retained.
Running SIMCA analysis is now possible using the Chemometrics tab. A cross-validation is initially run (the same settings as those used for PCA analysis are available). The SIMCA panel shows the resulting Model Table. Additional plots, Influence plots (for each class) and Cooman's plot (for each pair of classes), can be also obtained.
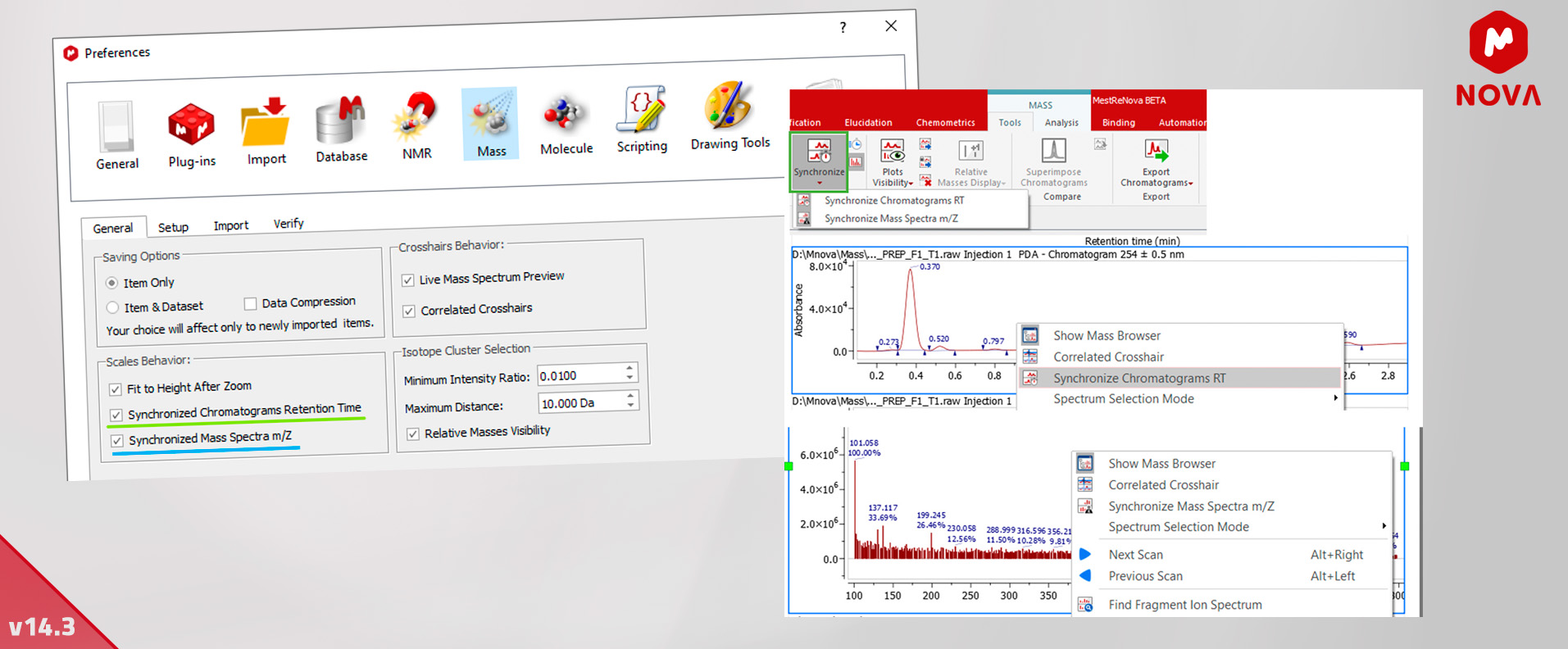
12. Improvements in Plot Scales - MSChrom
We have implemented the synchronization of the retention time for MS chromatograms and also added synchronization of m/z scale of mass spectra.
You can enable these features by default in Mnova > Preferences > Mass > General > Scales behavior and also control the synchronization using the button in menu Mass > Tools in the ribbon.
In addition, we have improved the way traces are synchronized when zooming in, even when traces don't contain data in a certain range, displaying an empty region.
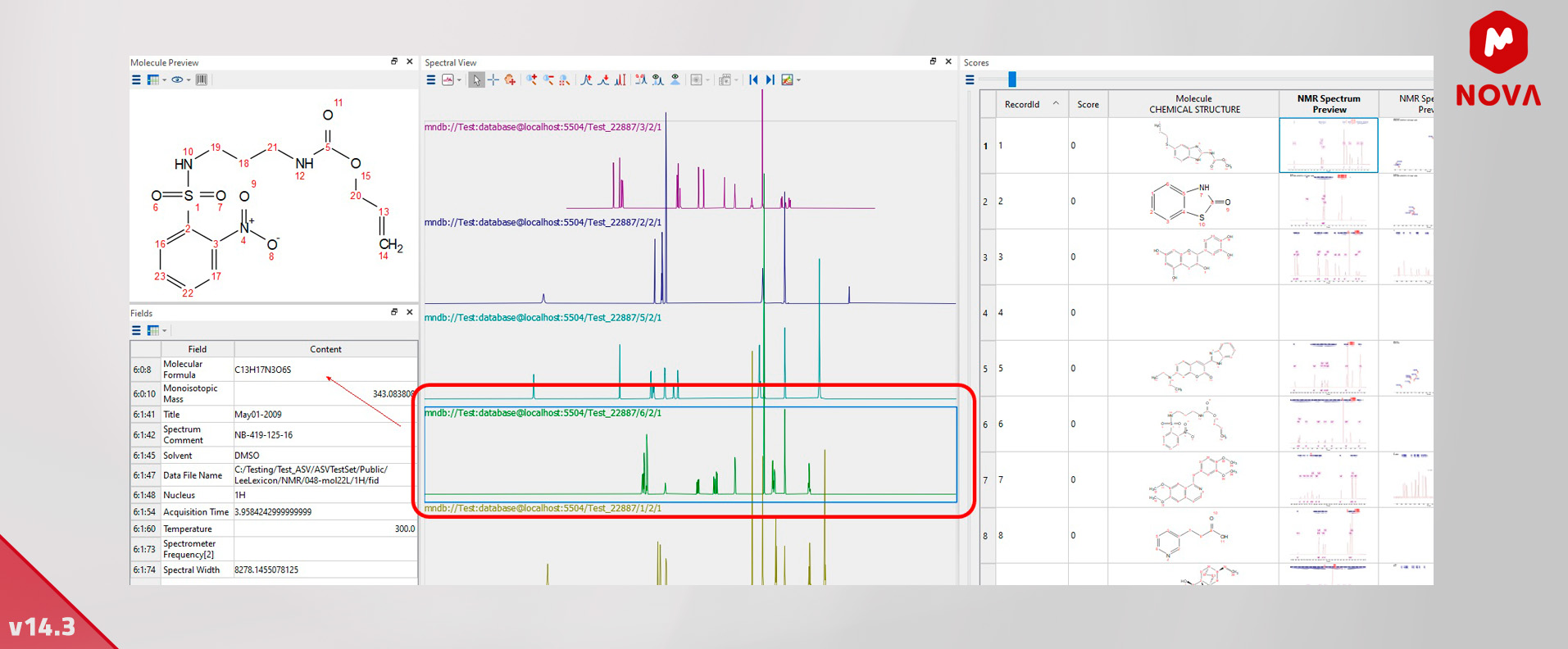
13. New Interactive Database Browser - Mnova DB
We have generated a new interactive database browser to help you with your database queries.
This tool contains a ‘spectral view’ widget to help you to interact with your single spectra or in a stacked mode view in Mnova DB without the need to copy your queries to Mnova itself.
Download Mnova 14.3
You can review our many other new and innovative features for all products and analysis types in our Changelog section.