Using qNMR
For this tutorial we will use a spectrum of acetanilide (acetanilide.mnova) with the applicable molecular structure. These are high quality data, well phased and the sample is very pure. The correct result is 20mM.
Generally, the workflow will be:
- Determine multiplets
- Run the script and select the qNMR variables
- The software will choose the multiplets to include
- Report
So first of all, we will load the 1H NMR spectrum and apply the necessary processing. (Depending on the level of accuracy required, you must pay attention to the number of data points used for the FT (typically 64K) and to apply a phase correction. Sometimes a baseline correction could be also needed. If necessary (e.g., solvent saturation was used), blind regions should be used to mask these spectral regions).
In the example shown below, we have just loaded the dataset (which you can download) and applied an 'automatic multiplet analysis':
The qNMR functionality is selected from the “Advanced” menu item. The software stores all settings in the registry, so you may not have to change things. But you will need to provide some information the first time you use this. This will be described in more detail in the manual. In this tutorial we will click on the ‘Settings’ button of the qNMR panel and then on the ‘Load Parameters’ button to load the ‘1_acetanilide.data’ file (which you have downloaded in the previous step)
After having clicked on the OK button, the multiplets will be found and (absolute) integrals found for their bounds. Having the correct number of nuclides is important and the software may do this better if the molecular structure is available and plotted.
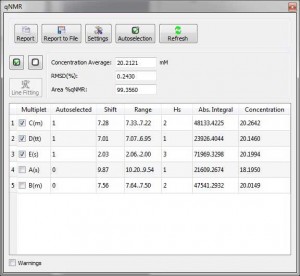
At this stage the qNMR process will be done automatically and the multiplets sorted in the table based on their perceived ranking as candidates for qNMR. The selected multiplets can be changed and the effect on the derived concentration. The concetration averaged calculated for this example is 20.2 mM.