Use Mnova to make high quality images for your publications
Mnova is a "WYSIWYG" application: what you see on the screen is what you will get when you produce files or hard copy of your prized data. But converting your spectral images into graphics for various purposes requires a little thought. There are a number of ways to produce spectra for export. The simplest is a screen capture, followed by pasting. Next you can create image files, and lastly you may want to make a PDF file - especially useful for laboratory notebooks and ELNs. A graphics guru,working in Adobe Creative Studio could undoubtedly write a more precise article, but this description is "by a chemist, for a chemist" using Mnova.
If you really want to get serious...
If you want to get serious with graphics then you need to learn about these terms, and constantly consider them:
- dots per inch (DPI)
- pixels per inch (PPI)
- Bitmap graphics
- Vector graphics
You can choose from specialized, expensive software such as Adobe's Photoshop Creative Studio, or Gimp, which is free - and many more. iDraw seems a reasonably-priced, capable vector drawing/editing option for Mac users. paint.net may be a good alternative. In this example I used "Recolor" in paint.net to do some basic editing. Note that this is a bitmap image with 96 DPI, which does not have fantastic quality.
Get the data acquisition right
If you do a poor job of data collection, then no amount of editing software can turn the "pig's ear" into a "silk purse". Having said that, there is considerable editing power in Mnova that can help a lot.
Common issues to consider are:
- poor shimming
- impurity levels
- using old, wet solvents, chipped NMR tubes, samples having solid particles, etc.
- poor signal-to-noise
- having too few data points
You may not have control over these, but when you do then get it right - in the spectrometer. For example, the number of data points can be easily controlled, and with huge disk storage space now the norm, there is no excuse to scrimp. If your lines are sharp then your acquisition time be at least a few seconds to avoid truncation artifacts. This applies especially to the direct dimension, and indirect dimension acquisition times of ca 100 ms will produce poor resolution and terrible wiggles. Remember that you can use non-uniform sampling (NUS) to improve digital resolution in the indirect dimension whilst minimizing the experiment time.
Data processing
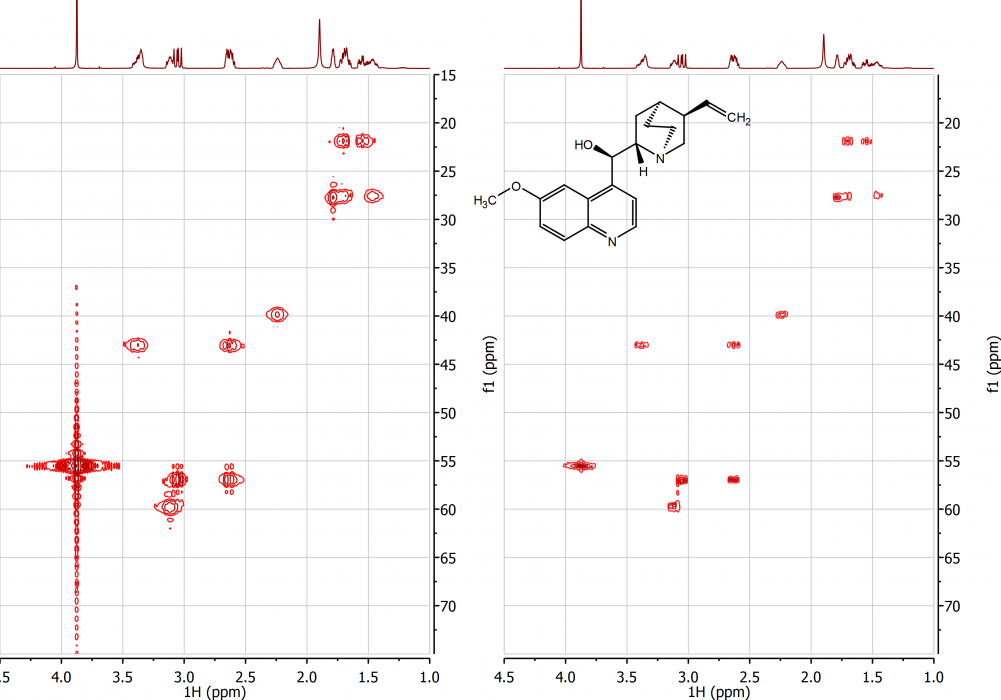
I stressed the need for good data acquisition, but data processing in Mnova is also your friend when you get back to your desk. If your FIDs are truncated they will produce ugly wiggles in the frequency domain spectrum. Use zero filling and as much line broadening apodization as you need to remove this from 1D spectra. With 2D spectra, forward linear prediction (LP) is a big help, and Mnova proprietary algorithms like "Reduce t1 noise" are very effective, too. Please see examples, below. I show the 2D HSQC spectrum of quinine, which is supplied with Mnova. On the LHS I used the 1K x 1K data points without apodization. On the RHS I show the effect of increasing the number of points to 4K in both dimensions, and performing LP. For apodization I used a sine-bell filter, shifted 45 degrees. t1 noise was reduced using the Mnova function.
2D HSQC showing the effect of processing
Getting the preferences right
What looks acceptable to you when viewing on the screen may not be best for printed copy or images. I generally find that the Mnova default line widths and font sizes are too thin/small - especially for output. As a rough guide I use a line width of 3.5 points, and font size of 12. I will use even thicker lines when they are straight, like chemical bonds, axes and borders - maybe 5 points. After changing your preferences, remember to press the "Set as Default" button at the bottom of the Preferences box. That way your customized values are always used. Remember, too, that preferences can be saved and read.
Don't forget that molecules can also have their preferences set to your liking. To do this, click with the right mouse button on the molecule object (green) grab handle, and a list will appear. At the bottom you will see "Preferences", and from there you can set attributes. My first attack is to thicken lines (ca 5 points) and change fonts (ca size 12). Again, you "Set as Default" your customization.
Data integrity
Mnova has some excellent tools to simplify data analysis. Baseline correction, solvent peak removal, for example, are often used to assist spectrum analysis. "Cuts" are useful in that they focus on/expand regions of interest. Unfortunately they can leave the cynical reader wondering what might be omitted - a nasty impurity? I would always show the full spectrum, and then perhaps some expansions that draw attention to regions of special interest. Many have expressed opinions about attempts in the literature to hide spikes, impurities, etc. by covering with white boxes and so forth. Others try to pass simulated spectra as real. DON'T DO IT! Organic Letters has studies this problem and has a clear data integrity policy.
Screen shots
You may use Print Screen in Windows, for example - or maybe something a bit more fancy like "Snagit" to copy the image to the clipboard. Either way you invariably get an image that has the screen resolution in terms of its ultimate quality. The dots per inch (dpi) for this may not be fantastic - usually 96 - and I therefore only advise this when you will be ultimately showing the image on the screen. For me this is the case with a Powerpoint presentation.
In the image, below, I used the exact same data as above. However, I reduced the screen resolution, and copied the file as a JPEG image (rather than PNG).
Publication quality
Many publications ask you provide high quality graphical images. In reality this often means that the DPI resolution should be 600 or better, and a vector image used. This is very easy to do using Mnova! This approach requires that you save a file for each figure. This is a good idea because these can included in any of your presentations - you can insert an image into Microsoft Office programs. And it may be easier to organize your images as separate files rather than looking for them in presentations.
Simply follow these easy steps:
1. Compose your image
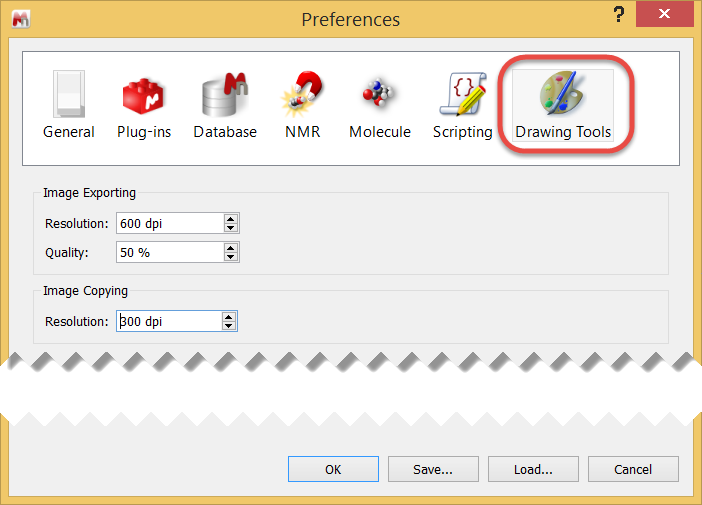
2. Under "Edit / Preferences / Drawing tools" set your image export resolution (e.g., 300 or 600 dpi)
3. In Mnova, select "File / Save As..." From the "Save as type" pull down menu choose a suitable graphics format. 4. Select the directory, give the file a name, and you are done! 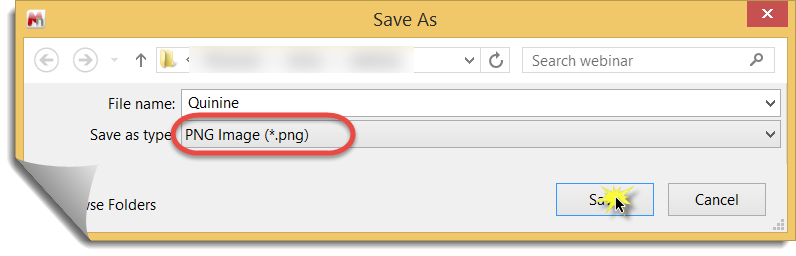
Conclusions
Mnova is a wonderful tool to process and analyze your spectroscopic data. Producing amazing copies of these for everything from Powerpoint presentations to files for publications is a quick, simple matter.